Ion Proton Sequencer ... Now that's a formidable name. And it's one that could be applied to just about any technology of the phantasmagorical sci-fi ilk. Was it that thing they used to beam Kirk, Spock, Bones and a hapless "Red Shirt" to the surface of every unknown planet? Negative. That was a transporter. Is it one of Dr. Thaddeus Venture's failed inventions from "The Venture Bros." series? Nope, you're thinking of the metasonic locator on that one.
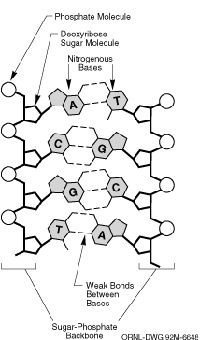
If the Ion Proton Sequencer were a piece of fiction -- which it's not -- it would be found in the medical lab of every deep-space exploratory starship; it would study new life forms and figure out just how many trips on the transporter would reduce your DNA's double helix to something that looks like a fireman's pole. But alas, the machine is a real and revolutionary medical research device.
Advertisement
It was the first in a new generation of sequencers that can decode an entire human genome quickly and inexpensively. Minus the time to prepare the DNA sample, processing only takes about a day. And the company says the test costs about $1,000.
Those two factors make this machine -- and those of its competitors, like Illumina and Pacific Biosciences -- a complete game changer for medical research and treatment. Until recently, genetic decoding was still rather rare due to the prohibitive costs and time involved. It could take days or weeks just to have partial DNA sequences read. And a full decoding of a person's genome cost hundreds of thousands of dollars just a few years ago [source: NOVA].
As a result, this sort of technology was usually reserved for the lab environment. Unless doctors or researchers were dealing with an outstanding medical case with widespread health ramifications, such genetic scrutiny usually wasn't practical.
One of those outstanding cases took place in 2011, when a new strain of E. coli broke out in Germany. While investigators were busy tracking down the source, researchers were looking at the makeup of this new bacterium. They used a predecessor of the Ion Proton Sequencer called the Personal Genome Machine (PGM) Sequencer. It's made by the same company, Life Technologies, and could read long portions of the organism's genetic blueprint in hours as opposed to weeks.
Researchers were able to compare that blueprint with previously decoded E. coli and determine that they were dealing with a new variant and understand some of its traits, such as toxicity and which antibiotics it might resist. Armed with that data, doctors stood a better chance at helping patients infected with the pathogen. Ultimately, more than 4,000 people were infected, and 50 known cases were killed by it [source: World Health Organization].
So, we know genetic sequencing is powerful stuff. It's already proven that it can help stop an outbreak. Let's learn about the developments that made the Ion Proton Sequencer possible and what it could do for future medicine.
Advertisement